-Search query
-Search result
Showing all 28 items for (author: petrovic & a)

EMDB-29172: 
Cryo-EM structure of Cryptococcus neoformans trehalose-6-phosphate synthase homotetramer in complex with uridine diphosphate and glucose-6-phosphate
Method: single particle / : Washington EJ, Brennan RG

PDB-8fhw: 
Cryo-EM structure of Cryptococcus neoformans trehalose-6-phosphate synthase homotetramer in complex with uridine diphosphate and glucose-6-phosphate
Method: single particle / : Washington EJ, Brennan RG

EMDB-16458: 
Electron cryo-tomography and subtomogram averaging of cytoplasmic lattice filaments from mammalian oocytes
Method: subtomogram averaging / : Petrovic A, Bauerlein FJB, Jentoft IMA, Schuh M, Fernandez-Busnadiego R

EMDB-16472: 
In situ cryo-electron tomogram of cytoplasmic lattice filaments from a mouse oocyte
Method: electron tomography / : Bauerlein FJB, Jentoft IMA, Petrovic A, Fernandez-Busnadiego R, Schuh M

PDB-7tbj: 
Composite structure of the human nuclear pore complex (NPC) symmetric core generated with a 12A cryo-ET map of the purified HeLa cell NPC
Method: subtomogram averaging / : Petrovic S, Samanta D, Perriches T, Bley CJ, Thierbach K, Brown B, Nie S, Mobbs GW, Stevens TA, Liu X, Tomaleri GP, Schaus L, Hoelz A

PDB-7tbl: 
Composite structure of the human nuclear pore complex (NPC) cytoplasmic face generated with a 12A cryo-ET map of the purified HeLa cell NPC
Method: subtomogram averaging / : Bley CJ, Nie S, Mobbs GW, Petrovic S, Gres AT, Liu X, Mukherjee S, Harvey S, Huber FM, Lin DH, Brown B, Tang AW, Rundlet EJ, Correia AR, Chen S, Regmi SG, Stevens TA, Jette CA, Dasso M, Patke A, Palazzo AF, Kossiakoff AA, Hoelz A

EMDB-24056: 
Single particle cryo-EM structure of the Chaetomium thermophilum Nup192-Nic96 complex (Nup192 residues 1-1756; Nic96 residues 240-301)
Method: single particle / : Petrovic S, Samanta D, Perriches T, Bley CJ, Thierbach K, Brown B, Nie S, Mobbs GW, Stevens TA, Liu X, Tomaleri GP, Schaus L, Hoelz A

EMDB-24057: 
Single particle cryo-EM structure of the Chaetomium thermophilum Nup192-Nic96-Nup53-Nup145N complex (Nup192 residues 1-1756; Nic96 residues 240-301; Nup53 31-67; Nup145N 616-683)
Method: single particle / : Petrovic S, Samanta D, Perriches T, Bley CJ, Thierbach K, Brown B, Nie S, Mobbs GW, Stevens TA, Liu X, Tomaleri GP, Schaus L, Hoelz A

EMDB-24058: 
Single particle cryo-EM structure of the Chaetomium thermophilum Nup188-Nic96 complex (Nup188 residues 1-1858; Nic96 residues 240-301)
Method: single particle / : Petrovic S, Samanta D, Perriches T, Bley CJ, Thierbach K, Brown B, Nie S, Mobbs GW, Stevens TA, Liu X, Tomaleri GP, Schaus L, Hoelz A

EMDB-24059: 
Single particle cryo-EM structure of the Chaetomium thermophilum Nup188-Nic96-Nup145N complex (Nup188 residues 1-1858; Nic96 residues 240-301; Nup145N residues 640-732)
Method: single particle / : Petrovic S, Samanta D, Perriches T, Bley CJ, Thierbach K, Brown B, Nie S, Mobbs GW, Stevens TA, Liu X, Tomaleri GP, Schaus L, Hoelz A

PDB-7mvu: 
Single particle cryo-EM structure of the Chaetomium thermophilum Nup192-Nic96 complex (Nup192 residues 1-1756; Nic96 residues 240-301)
Method: single particle / : Petrovic S, Samanta D, Perriches T, Bley CJ, Thierbach K, Brown B, Nie S, Mobbs GW, Stevens TA, Liu X, Tomaleri GP, Schaus L, Hoelz A

PDB-7mvv: 
Single particle cryo-EM structure of the Chaetomium thermophilum Nup192-Nic96-Nup53-Nup145N complex (Nup192 residues 1-1756; Nic96 residues 240-301; Nup53 31-67; Nup145N 616-683)
Method: single particle / : Petrovic S, Samanta D, Perriches T, Bley CJ, Thierbach K, Brown B, Nie S, Mobbs GW, Stevens TA, Liu X, Tomaleri GP, Schaus L, Hoelz A

PDB-7mvy: 
Single particle cryo-EM structure of the Chaetomium thermophilum Nup188-Nic96 complex (Nup188 residues 1-1858; Nic96 residues 240-301)
Method: single particle / : Petrovic S, Samanta D, Perriches T, Bley CJ, Thierbach K, Brown B, Nie S, Mobbs GW, Stevens TA, Liu X, Tomaleri GP, Schaus L, Hoelz A

PDB-7mvz: 
Single particle cryo-EM structure of the Chaetomium thermophilum Nup188-Nic96-Nup145N complex (Nup188 residues 1-1858; Nic96 residues 240-301; Nup145N residues 640-732)
Method: single particle / : Petrovic S, Samanta D, Perriches T, Bley CJ, Thierbach K, Brown B, Nie S, Mobbs GW, Stevens TA, Liu X, Tomaleri GP, Schaus L, Hoelz A

PDB-7tbi: 
Composite structure of the S. cerevisiae nuclear pore complex (NPC)
Method: subtomogram averaging / : Petrovic S, Samanta D, Perriches T, Bley CJ, Thierbach K, Brown B, Nie S, Mobbs GW, Stevens TA, Liu X, Tomaleri GP, Schaus L, Hoelz A

PDB-7tbk: 
Composite structure of the dilated human nuclear pore complex (NPC) symmetric core generated with a 37A in situ cryo-ET map of CD4+ T cell NPC
Method: subtomogram averaging / : Petrovic S, Samanta D, Perriches T, Bley CJ, Thierbach K, Brown B, Nie S, Mobbs GW, Stevens TA, Liu X, Tomaleri GP, Schaus L, Hoelz A

PDB-7tbm: 
Composite structure of the dilated human nuclear pore complex (NPC) generated with a 37A in situ cryo-ET map of CD4+ T cell NPC
Method: subtomogram averaging / : Bley CJ, Nie S, Mobbs GW, Petrovic S, Gres AT, Liu X, Mukherjee S, Harvey S, Huber FM, Lin DH, Brown B, Tang AW, Rundlet EJ, Correia AR, Chen S, Regmi SG, Stevens TA, Jette CA, Dasso M, Patke A, Palazzo AF, Kossiakoff AA, Hoelz A

EMDB-25474: 
CryoEM structure of the N-terminal-deleted Rix7 AAA-ATPase
Method: single particle / : Kocaman S, Stanley RE, Lo YH, Krahn J, Dandey VP, Sobhany M, Petrovich M, Williams JG, Deterding LJ, Borgnia MJ, Etigunta S

EMDB-25582: 
CryoEM structure of the crosslinked Rix7 AAA-ATPase
Method: single particle / : Kocaman S, Stanley RE, Lo YH, Krahn J, Dandey VP, Sobhany M, Petrovich M, Williams JG, Deterding LJ, Borgnia MJ, Etigunta S

EMDB-25659: 
CryoEM structure of the Rix7 D2 Walker B mutant
Method: single particle / : Lo YH, Krahn J

PDB-7swl: 
CryoEM structure of the N-terminal-deleted Rix7 AAA-ATPase
Method: single particle / : Kocaman S, Stanley RE, Lo YH, Krahn J, Dandey VP, Sobhany M, Petrovich M, Williams JG, Deterding LJ, Borgnia MJ, Etigunta S

PDB-7t0v: 
CryoEM structure of the crosslinked Rix7 AAA-ATPase
Method: single particle / : Kocaman S, Stanley RE, Lo YH, Krahn J, Dandey VP, Sobhany M, Petrovich M, Williams JG, Deterding LJ, Borgnia MJ, Etigunta S

PDB-7t3i: 
CryoEM structure of the Rix7 D2 Walker B mutant
Method: single particle / : Kocaman S, Stanley RE

EMDB-7326: 
Cryo-EM structure of human kinetochore protein CENP-N with the centromeric nucleosome containing CENP-A
Method: single particle / : Zhou K, Pentakota S, Vetter IR

PDB-6c0w: 
Cryo-EM structure of human kinetochore protein CENP-N with the centromeric nucleosome containing CENP-A
Method: single particle / : Zhou K, Pentakota S, Vetter IR, Morgan GP, Petrovic A, Musacchio A, Luger K

EMDB-4103: 
Structure of the human Rod-Zw10-Zwilch (RZZ) complex
Method: single particle / : Mosalaganti S, Keller J

EMDB-4104: 
Negative stain reconstruction of the ROD(1-1250):Zwilch:ZW10
Method: single particle / : Mosalaganti S
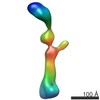
EMDB-2549: 
3D structure of the KMN network
Method: single particle / : Petrovic A, Mosalaganti S, Keller J, Mattiuzzo M, Overlack K, Krenn V, De Antoni A, Wohlgemuth S, Cecatiello V, Pasqualato S, Raunser S, Musacchio A
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
